https://github.com/compomics/jsparklines
Sparklines for java tables
https://github.com/compomics/jsparklines
sparkline visualization
Last synced: about 2 months ago
JSON representation
Sparklines for java tables
- Host: GitHub
- URL: https://github.com/compomics/jsparklines
- Owner: CompOmics
- Created: 2015-07-02T13:06:43.000Z (almost 11 years ago)
- Default Branch: master
- Last Pushed: 2023-03-01T14:03:41.000Z (about 3 years ago)
- Last Synced: 2025-09-10T05:11:34.154Z (7 months ago)
- Topics: sparkline, visualization
- Language: Java
- Homepage: http://compomics.github.io/projects/jsparklines.html
- Size: 4.26 MB
- Stars: 9
- Watchers: 11
- Forks: 2
- Open Issues: 2
-
Metadata Files:
- Readme: README.md
- License: LICENSE-2.0.txt
Awesome Lists containing this project
README
# JSparklines
JSparklines makes it straightforward to visualize numbers in Java tables by the use of [sparklines](http://en.wikipedia.org/wiki/Sparklines). All that is needed is a couple of lines of code.
The charts are created using [JFreeChart](http://www.jfree.org/jfreechart) and added to the table columns using custom [TableCellRenderers](http://download.oracle.com/javase/tutorial/uiswing/components/table.html#renderer).
Supports more than **30 types of charts/renderers**, including bar chart, line charts, stacked bar charts, bar charts with error bars, pie charts, scatter plots, interval charts, area charts, heat maps and box plots.
If you like JSparklines you may also want to check out [JSparklines Factory](https://github.com/sing-group/GC4S/tree/master/gc4s-jsparklines-factory).
---
## JSparklines Publication:
* [Barsnes et al: Proteomics. 2014. (doi: 10.1002/pmic.201400356)](http://www.ncbi.nlm.nih.gov/pubmed/25422159).
* If you use JSparklines as part of a publication, please include this reference.
---
| | | | |
| :------------------------- | :---------------: | :--: | :--: |
| [](https://genesis.ugent.be/maven2/no/uib/jsparklines/1.1.2/jsparklines-1.1.2.zip) | *v1.1.2 - All platforms* | [ReleaseNotes](https://github.com/compomics/jsparklines/wiki/ReleaseNotes) | [JavaDoc](http://compomics.github.io/jsparklines/javadoc) |
---
## How to use JSparklines
See the [How to use JSparklines](https://github.com/compomics/jsparklines/wiki/Tutorial) wiki page for code examples. JSparklines is also available as a [Maven dependency](https://github.com/compomics/jsparklines/wiki/Tutorial#maven-dependency).
_If you have questions or would like to contribute to the **JSparklines** project, please contact the developers._
---
## Examples
| | |
| :--: | :--: |
| [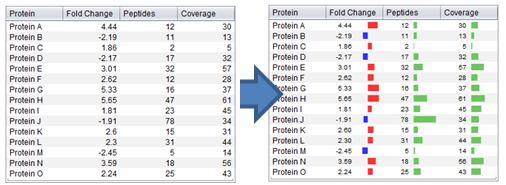](https://github.com/compomics/jsparklines/wiki/images/JSparklinesSample.png) | [](https://github.com/compomics/jsparklines/wiki/images/JSparklinesDemo2.png) |
_(Click on a figure to see the full size version)_
---
## Projects using JSparklines ##
| Project | Description | Publication |
|:-----------------|:----------------|:----------------|
| [PeptideShaker](http://compomics.github.io/projects/peptide-shaker.html) | _interpretation of proteomics identification results_|_[Vaudel et al: Nature Biotechnol. 2015 Jan;33(1):22-24.](http://www.nature.com/nbt/journal/v33/n1/full/nbt.3109.html)_|
| [DeNovoGUI](http://compomics.github.io/projects/denovogui.html) | _de novo sequencing of tandem mass spectra_|_[Muth at al: J Proteome Res. 2014 Feb 7;13(2):1143-6.](http://www.ncbi.nlm.nih.gov/pubmed/24295440)_|
| [SearchGUI](http://compomics.github.io/projects/searchgui.html) | _graphical user interface for proteomics identification search engines_|_[Barsnes et al: J Proteome Res. 2018 (in press)](https://www.ncbi.nlm.nih.gov/pubmed/29774740)_|
| [thermo-msf-parser](http://compomics.github.io/projects/thermo-msf-parser.html) | _parser and viewer for thermo msf files_|_[Colaert et al: J Proteome Res. 2011;10(8):3840-3.](http://www.ncbi.nlm.nih.gov/pubmed/21714566)_|
| [Fragmentation Analyzer](http://compomics.github.io/projects/fragmentation-analyzer.html) | _analyzing MS/MS fragmentation data_|_[Barsnes et al: Proteomics 2010;10(5):1087-90.](http://www.ncbi.nlm.nih.gov/pubmed/20049869)_|
| [MetaProteomeAnalyzer](https://github.com/compomics/meta-proteome-analyzer) | _analyzing meta-proteomics data_|_[Muth et al: J Proteome Res. 2015 Mar 6;14(3):1557-65.](http://www.ncbi.nlm.nih.gov/pubmed/25660940)_|
| [proteocloud](http://code.google.com/p/proteocloud) | _proteomics cloud computing pipeline_|_[Muth et al: J Proteomics. 2013 Jan 8. pii: S1874-3919(13)00013-4.](http://www.ncbi.nlm.nih.gov/pubmed/23305951)_|
| [S2P](http://www.sing-group.org/s2p) | _processing of 2D-gel and MALDI-based mass spectrometry protein data_|_[López-Fernández et al: Comput Methods Programs Biomed. 2018;Volume 155, pp. 1-9](https://www.sciencedirect.com/science/article/pii/S0169260717310702)_|
| [DEWE](http://www.sing-group.org/dewe) | _differential expression analysis of RNA-Seq data_| |
_Are you using JSparklines and would like your project listed here? Contact the developers of JSparklines._
---