https://github.com/deeplearnphysics/spine
Scalable Particle Imaging with Neural Embeddings
https://github.com/deeplearnphysics/spine
machine-learning neutrino-physics particle-imaging reconstruction
Last synced: about 2 months ago
JSON representation
Scalable Particle Imaging with Neural Embeddings
- Host: GitHub
- URL: https://github.com/deeplearnphysics/spine
- Owner: DeepLearnPhysics
- License: mit
- Created: 2024-03-27T23:23:14.000Z (about 2 years ago)
- Default Branch: main
- Last Pushed: 2026-04-05T21:14:46.000Z (about 2 months ago)
- Last Synced: 2026-04-05T23:21:25.779Z (about 2 months ago)
- Topics: machine-learning, neutrino-physics, particle-imaging, reconstruction
- Language: Python
- Homepage:
- Size: 8.45 MB
- Stars: 5
- Watchers: 3
- Forks: 18
- Open Issues: 13
-
Metadata Files:
- Readme: README.md
- Changelog: CHANGELOG.md
- Contributing: CONTRIBUTING.md
- License: LICENSE
Awesome Lists containing this project
README
[](https://github.com/DeepLearnPhysics/spine/actions/workflows/ci.yml)
[](https://codecov.io/gh/DeepLearnPhysics/spine)
[](https://spine.readthedocs.io/latest/)
[](https://badge.fury.io/py/spine)
[](https://pypi.org/project/spine/)
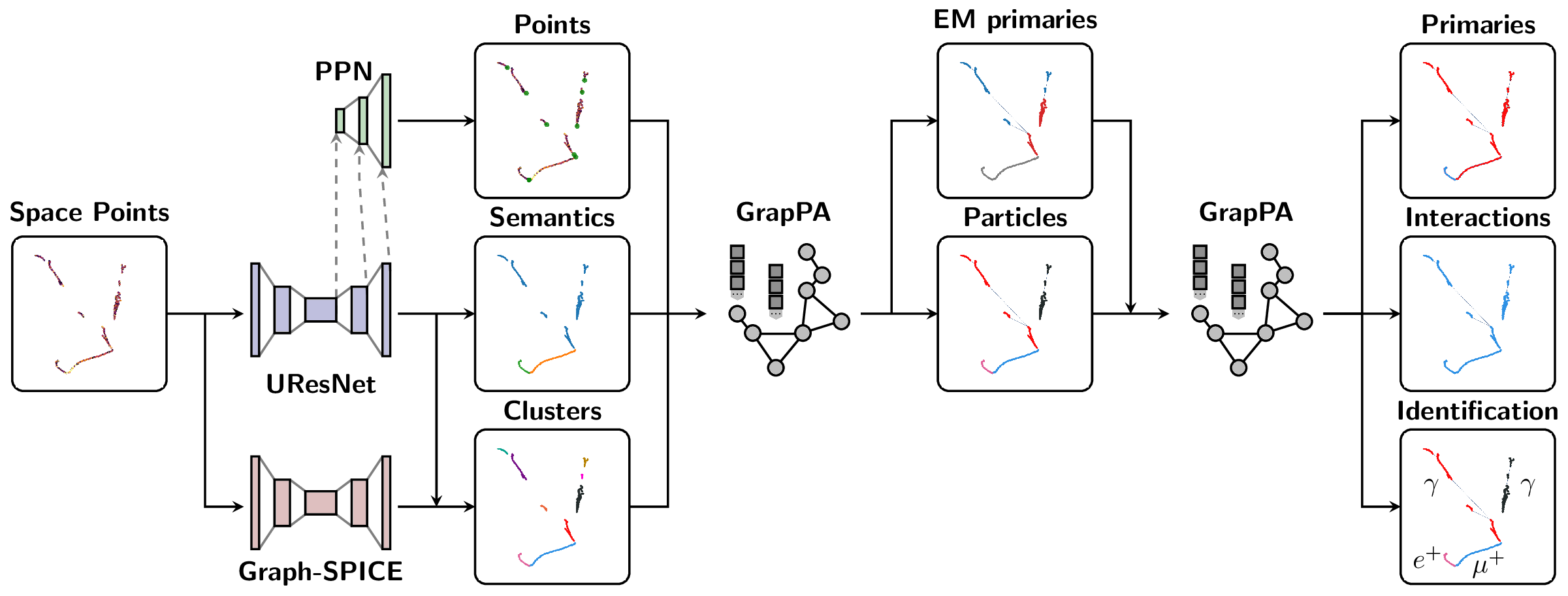
The Scalable Particle Imaging with Neural Embeddings (SPINE) package leverages state-of-the-art Machine Learning (ML) algorithms -- in particular Deep Neural Networks (DNNs) -- to reconstruct particle imaging detector data. This package was primarily developed for Liquid Argon Time-Projection Chamber (LArTPC) data and relies on Convolutional Neural Networks (CNNs) for pixel-level feature extraction and Graph Neural Networks (GNNs) for superstructure formation. The schematic below breaks down the full end-to-end reconstruction flow.
## Installation
SPINE is now available on PyPI with flexible installation options to suit different needs:
### Quick Start (Recommended)
For data analysis and visualization without machine learning:
```bash
pip install spine[all]
```
### Installation Options
**1. Core Package (minimal dependencies)**
```bash
# Essential dependencies: numpy, scipy, pandas, PyYAML, h5py, numba
pip install spine
```
**2. With Visualization Tools**
```bash
# Adds plotly, matplotlib, seaborn for data visualization
pip install spine[viz]
```
**3. Development Environment**
```bash
# Adds testing, formatting, and documentation tools
pip install spine[dev]
```
**4. Everything (except PyTorch)**
```bash
# All optional dependencies (visualization + development tools)
pip install spine[all]
```
### PyTorch ecosystem
#### Option 1: Container Approach (Recommended)
The easiest way to get a working PyTorch environment with LArCV support:
```bash
# Pull the SPINE-compatible container with complete PyTorch ecosystem + LArCV
singularity pull spine.sif docker://deeplearnphysics/larcv2:ub2204-cu121-torch251-larndsim
# Install SPINE in the container
singularity exec spine.sif pip install spine[all]
# Run your analysis
singularity exec spine.sif spine --config your_config.cfg --source data.h5
```
> This container includes: PyTorch 2.5.1, CUDA 12.1, torch-geometric, torch-scatter, torch-cluster, MinkowskiEngine, and **LArCV2**.
#### Option 2: Manual Installation** (advanced users):
```bash
# Step 1: Install PyTorch with CUDA
pip install torch --index-url https://download.pytorch.org/whl/cu118
# Step 2: Install ecosystem packages (critical order)
pip install --no-build-isolation torch-scatter torch-cluster torch-geometric MinkowskiEngine
# Step 3: Install SPINE
pip install spine[all]
```
> **� Why separate?** The PyTorch ecosystem (torch, torch-geometric, torch-scatter, torch-cluster, MinkowskiEngine) forms an interdependent group requiring exact version compatibility and complex compilation. Installing them together ensures compatibility.
### LArCV2
#### Option 1: Use the container (recommended)*
```bash
# LArCV2 is pre-installed in the DeepLearnPhysics container
singularity pull spine.sif docker://deeplearnphysics/larcv2:ub2204-cu121-torch251-larndsim
```
#### Option 2: Build from source*
```bash
# Clone and build the latest LArCV2
git clone https://github.com/DeepLearnPhysics/larcv2.git
cd larcv2
# Follow build instructions in the repository
```
> **Note**: Avoid conda-forge larcv packages as they may be outdated. Use the container or build from the official source.
### Development Installation
For developers who want to work with the source code:
```bash
git clone https://github.com/DeepLearnPhysics/spine.git
cd spine
pip install -e .[dev]
```
#### Quick Development Testing (No Installation)
For rapid development and testing without reinstalling the package:
```bash
# Clone the repository
git clone https://github.com/DeepLearnPhysics/spine.git
cd spine
# Install only the dependencies (not the package itself)
# Or alternatively simple run the commands inside the above container
pip install numpy scipy pandas pyyaml h5py numba psutil
# Run directly from source
python src/spine/bin/run.py --config config/train_uresnet.cfg --source /path/to/data.h5
# Or make it executable and run directly
chmod +x src/spine/bin/run.py
./src/spine/bin/run.py --config your_config.cfg --source data.h5
```
> **💡 Development Tip**: This approach lets you test code changes immediately without reinstalling. Perfect for rapid iteration during development.
To build and test packages locally:
```bash
# Build the package
./build_packages.sh
# Install locally built package
pip install dist/spine-*.whl[all]
```
## Usage
### Command Line Interface
**Option 1: After installation, use the `spine` command:**
```bash
# Run training/inference/analysis
spine --config config/train_uresnet.cfg --source /path/to/data.h5
```
**Option 2: Run directly from source (development):**
```bash
# From the spine repository directory
python src/spine/bin/run.py --config config/train_uresnet.cfg --source /path/to/data.h5
```
### Python API
Basic example:
```python
# Necessary imports
import yaml
from spine.driver import Driver
# Load configuration file
cfg_path = 'config/train_uresnet.cfg' # or your config file
with open(cfg_path, 'r') as f:
cfg = yaml.safe_load(f)
# Initialize driver class
driver = Driver(cfg)
# Execute model following the configuration regimen
driver.run()
```
* Documentation is available at https://spine.readthedocs.io/latest/.
* Tutorials and examples can be found in the documentation.
### Example Configuration Files
Example configurations are available in the `config` folder:
| Configuration name | Model |
| ------------------------------|----------------|
| `train_uresnet.cfg` | UResNet alone |
| `train_uresnet_ppn.cfg` | UResNet + PPN |
| `train_graph_spice.cfg` | GraphSpice |
| `train_grappa_shower.cfg` | GrapPA for shower fragments clustering |
| `train_grappa_track.cfg` | GrapPA for track fragments clustering |
| `train_grappa_inter.cfg` | GrapPA for interaction clustering |
To switch from training to inference mode, set `trainval.train: False` in your configuration file.
Key configuration parameters you may want to modify:
* `batch_size` - batch size for training/inference
* `weight_prefix` - directory to save model checkpoints
* `log_dir` - directory to save training logs
* `iterations` - number of training iterations
* `model_path` - path to checkpoint to load (optional)
* `train` - boolean flag for training vs inference mode
* `gpus` - GPU IDs to use (leave empty '' for CPU)
For more information on storing analysis outputs and running custom analysis scripts, see the documentation on `outputs` (formatters) and `analysis` (scripts) configurations.
### Running A Configuration File
Basic usage with the `spine` command:
```bash
# Run training/inference directly
spine --config config/train_uresnet.cfg --source /path/to/data.h5
# Or run in background with logging
nohup spine --config config/train_uresnet.cfg --source /path/to/data.h5 > log_uresnet.txt 2>&1 &
```
You can load a configuration file into a Python dictionary using:
```python
import yaml
# Load configuration file
with open('config/train_uresnet.cfg', 'r') as f:
cfg = yaml.safe_load(f)
```
### Reading a Log
A quick example of how to read a training log, and plot something
```python
import pandas as pd
import matplotlib.pyplot as plt
fname = 'path/to/log.csv'
df = pd.read_csv(fname)
# plot moving average of accuracy over 10 iterations
df.accuracy.rolling(10, min_periods=1).mean().plot()
plt.ylabel("accuracy")
plt.xlabel("iteration")
plt.title("moving average of accuracy")
plt.show()
# list all column names
print(df.columns.values)
```
### Recording network output or running analysis
Documentation for analysis tools and output formatting is available in the main documentation at https://spine.readthedocs.io/latest/.
## Repository Structure
* `bin` contains utility scripts for data processing
* `config` has example configuration files
* `docs` contains documentation source files
* `src/spine` contains the main package code
* `test` contains unit tests using pytest
Please consult the documentation for detailed information about each component.
## Testing and Coverage
### Running Tests
The SPINE package includes comprehensive unit tests using pytest:
```bash
# Run all tests
pytest
# Run tests for a specific module
pytest test/test_data/
# Run with verbose output
pytest -v
```
### Checking Test Coverage
Test coverage tracking helps ensure code quality and identify untested areas. Coverage reports are automatically generated in our CI pipeline and uploaded to [Codecov](https://codecov.io/gh/DeepLearnPhysics/spine).
To check coverage locally:
```bash
# Run the coverage script (generates terminal, HTML, and XML reports)
./bin/coverage.sh
# Or run pytest with coverage flags directly
pytest --cov=spine --cov-report=term --cov-report=html
# View the HTML report
open htmlcov/index.html
```
The coverage configuration is defined in `pyproject.toml` under `[tool.coverage.run]` and `[tool.coverage.report]`.
## Contributing
Before you start contributing to the code, please see the [contribution guidelines](CONTRIBUTING.md).
### Adding a new model
The SPINE framework is designed to be extensible. To add a new model:
1. **Data Loading**: Parsers exist for various sparse tensor and particle outputs in `spine.io.core.parse`. If you need fundamentally different data formats, you may need to add new parsers or collation functions.
2. **Model Implementation**: Add your model to the `spine.model` package. Include your model in the factory dictionary in `spine.model.factories` so it can be found by the configuration system.
3. **Configuration**: Create a configuration file in the `config/` folder that specifies your model architecture and training parameters.
Once these steps are complete, you should be able to train your model using the standard SPINE workflow.