https://github.com/dewberryants/asciiMol
Curses based ASCII molecule viewer for terminals.
https://github.com/dewberryants/asciiMol
ascii cheminformatics chemistry molecules visualization
Last synced: 3 months ago
JSON representation
Curses based ASCII molecule viewer for terminals.
- Host: GitHub
- URL: https://github.com/dewberryants/asciiMol
- Owner: dewberryants
- License: bsd-2-clause
- Created: 2020-09-03T12:48:52.000Z (about 5 years ago)
- Default Branch: master
- Last Pushed: 2025-02-17T08:08:58.000Z (8 months ago)
- Last Synced: 2025-06-02T06:07:29.546Z (4 months ago)
- Topics: ascii, cheminformatics, chemistry, molecules, visualization
- Language: Python
- Homepage:
- Size: 760 KB
- Stars: 381
- Watchers: 5
- Forks: 12
- Open Issues: 0
-
Metadata Files:
- Readme: README.md
- License: LICENSE
Awesome Lists containing this project
README
# asciiMOL
[](https://badge.fury.io/py/asciimol)
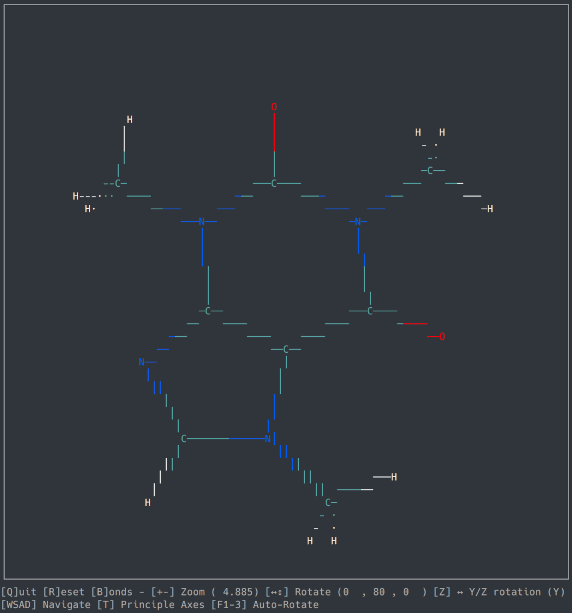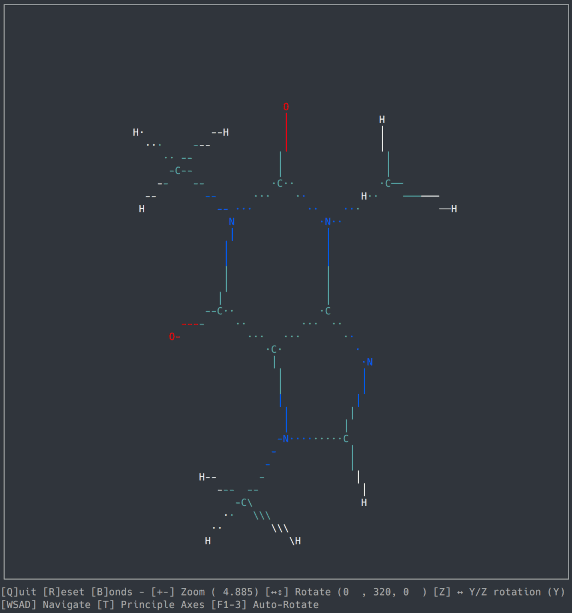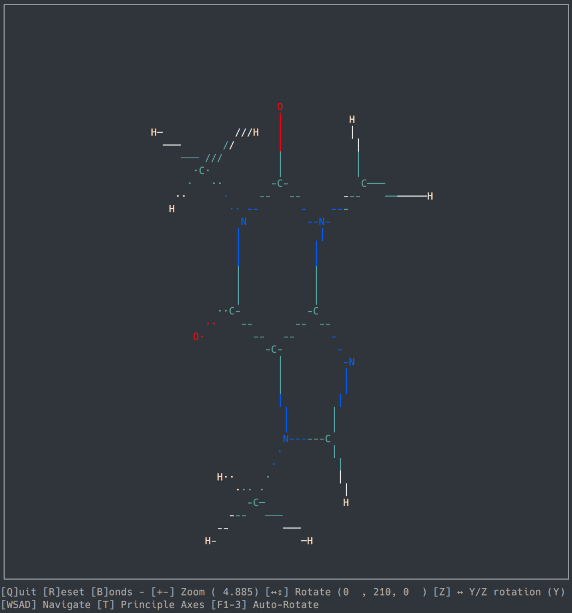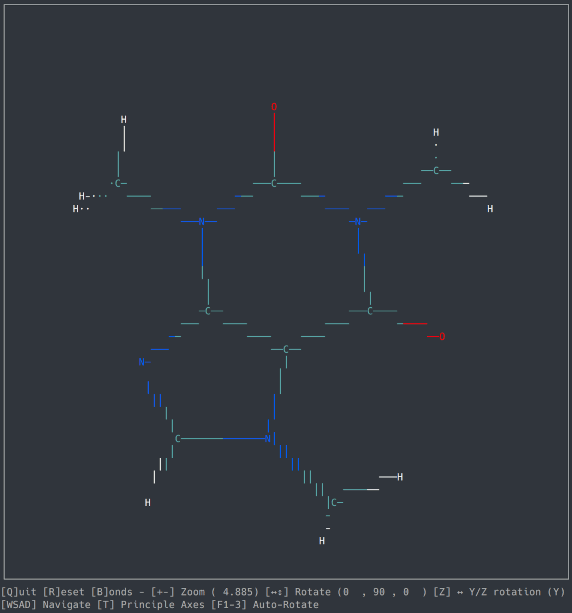
A basic molecule viewer written in Python, using the ncurses library. Works on any compatible terminal (even on Windows using
windows-curses).
Features:
* Opening default cartesian .xyz files
* Orthographic view
* Navigation
* Zoom, Rotation, Auto-Rotation
* Bond detection and display
* Support for simple .xyz trajectories
* Optional integration of ASE and RDKit pypi packages for more formats and SMILES
## Installation
```sh
pip install asciimol
```
(Note: pip will install a run script in $HOME/.local/bin/ if you do not install with root permissions, so make sure this
directory is part of your $PATH.)
You can also run
```sh
pip install asciimol[formats,smiles]
```
to automatically install ASE for formats and RDKit for smiles.