https://github.com/jb-sharp/medialaxis3d
Python implementation of 3D Medial Axis Transform for skeletonization.
https://github.com/jb-sharp/medialaxis3d
3d mat skeletonization
Last synced: 3 months ago
JSON representation
Python implementation of 3D Medial Axis Transform for skeletonization.
- Host: GitHub
- URL: https://github.com/jb-sharp/medialaxis3d
- Owner: jb-sharp
- Created: 2025-08-01T15:54:39.000Z (9 months ago)
- Default Branch: main
- Last Pushed: 2025-08-08T14:54:40.000Z (9 months ago)
- Last Synced: 2025-09-03T02:41:26.856Z (8 months ago)
- Topics: 3d, mat, skeletonization
- Language: Cython
- Homepage:
- Size: 4.1 MB
- Stars: 0
- Watchers: 0
- Forks: 0
- Open Issues: 0
-
Metadata Files:
- Readme: README.md
Awesome Lists containing this project
README
# medialaxis3d
This package extends the [scikit-image](https://scikit-image.org/) function [medial_axis](https://scikit-image.org/docs/stable/api/skimage.morphology.html#skimage.morphology.medial_axis)
to the 3D case.
## Install
```bash
pip install medialaxis3d
```
### Dependencies
Automatically installed with `pip`:
- `numpy`
- `scipy`
- `cython`
Optional only for visualization
- `napari`
## Documentation
WIP
## Quickstart
Use it without returning the medial distance.
```Python
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
>>> blob_size_fraction = 0.2,
>>> n_dim = 3,
>>> volume_fraction = 0.6,
>>> rng = rng)
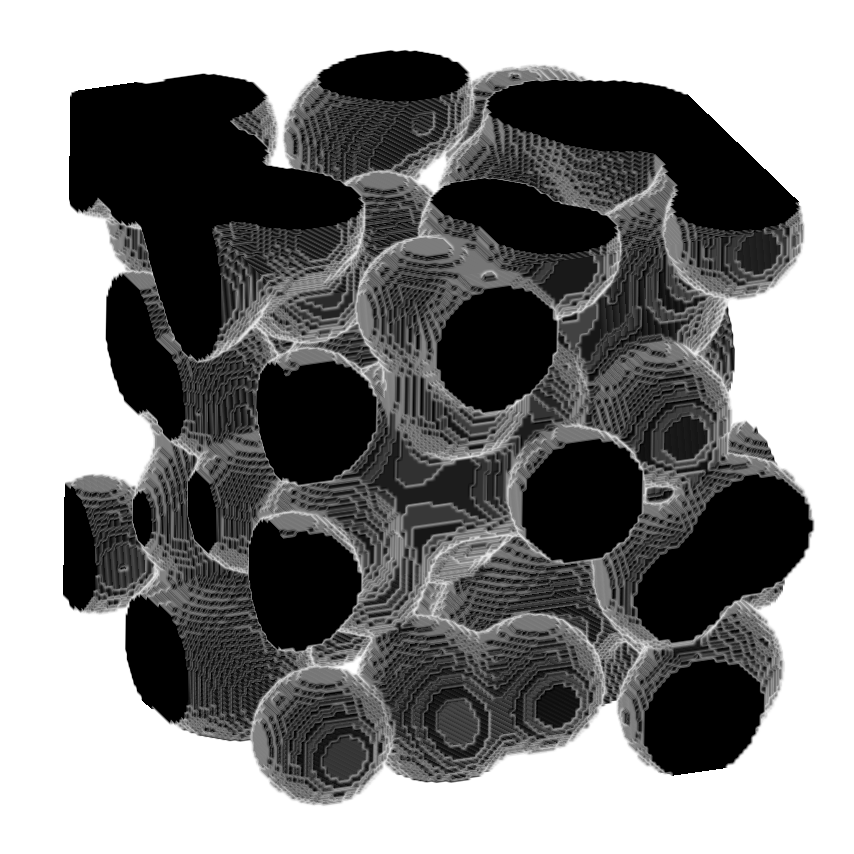
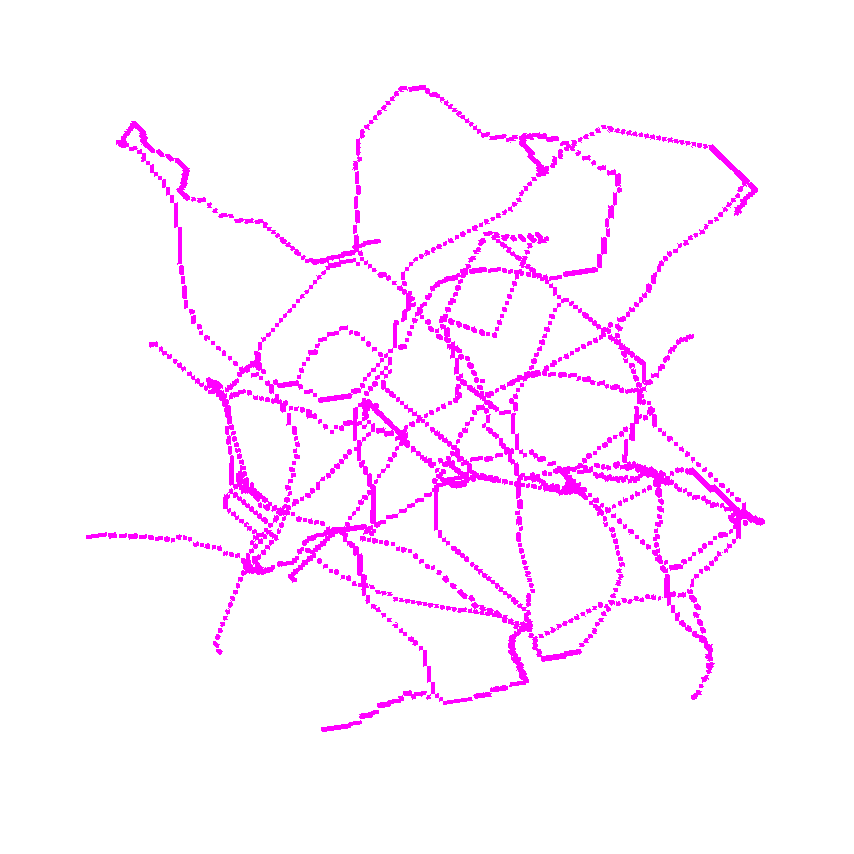
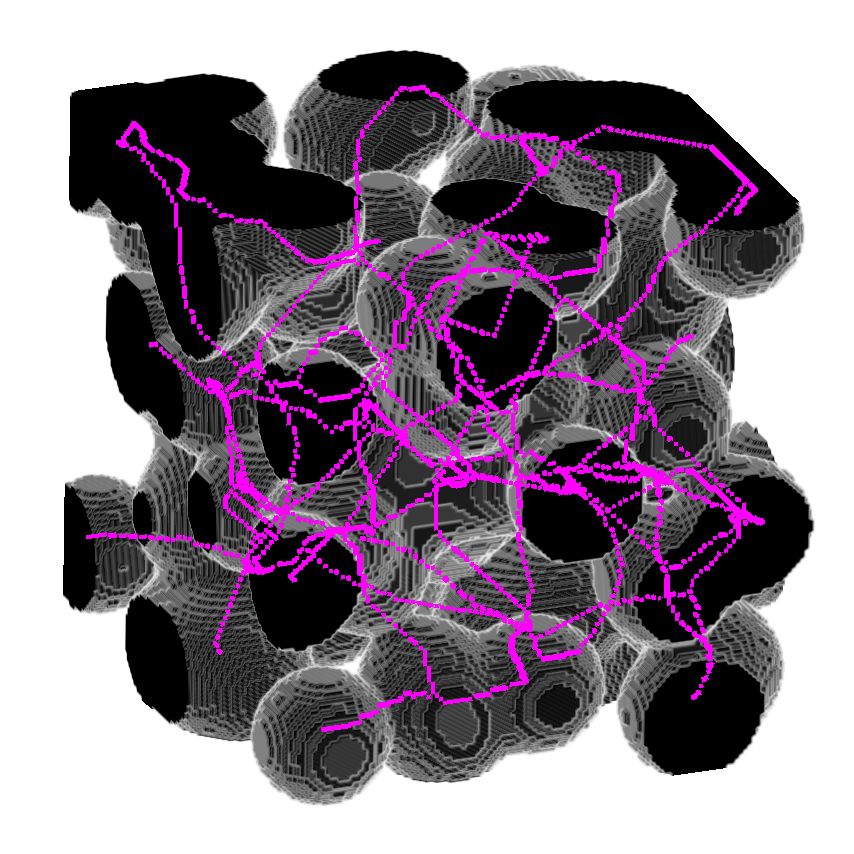
>>> skeleton = medialaxis3d.medial_axis_3d(image,
>>> return_distance = False,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton,
>>> interpolation3d = "nearest",
>>> colormap = "magenta",
>>> scale = [1, 1, 1])
>>> napari.run()
```

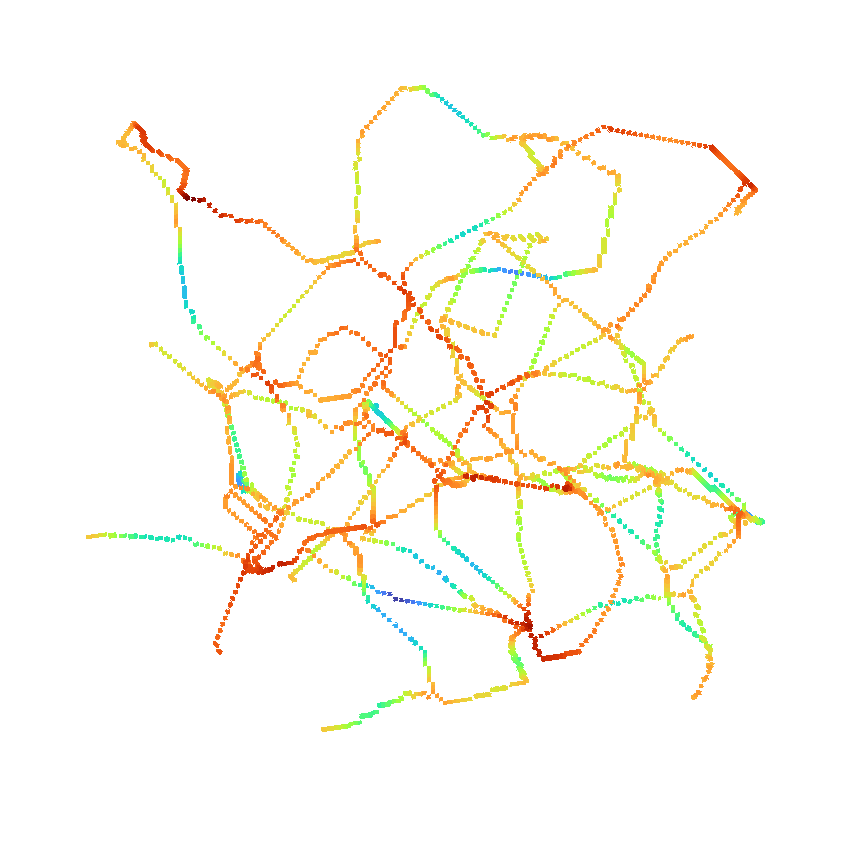
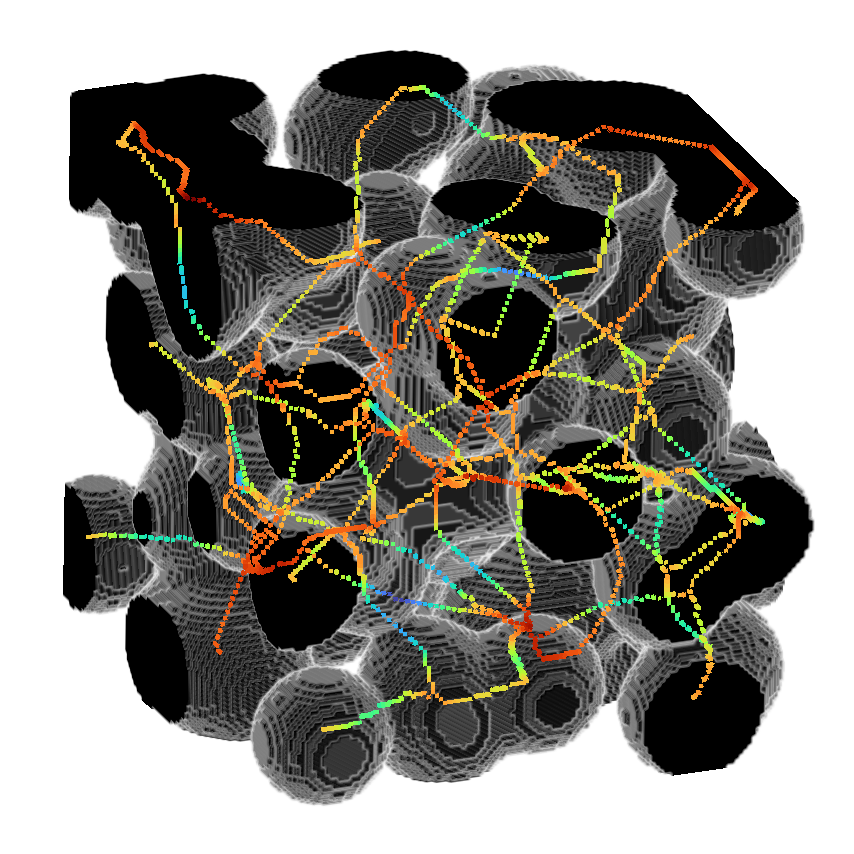
or use it to return the distance as well.
```Python
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
blob_size_fraction = 0.2,
n_dim = 3,
volume_fraction = 0.6,
rng = rng)
>>> skeleton, distance = medialaxis3d.medial_axis_3d(image,
>>> return_distance = True,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton*distance,
>>> interpolation3d = "nearest",
>>> colormap = "turbo",
>>> scale = [1, 1, 1])
>>> napari.run()
```