https://github.com/nstarman/mvgkde
MultiVariate Gaussian Kernel Density Estimation
https://github.com/nstarman/mvgkde
kernel-density-estimation multivariate-analysis
Last synced: 9 months ago
JSON representation
MultiVariate Gaussian Kernel Density Estimation
- Host: GitHub
- URL: https://github.com/nstarman/mvgkde
- Owner: nstarman
- License: apache-2.0
- Created: 2024-10-23T18:44:56.000Z (over 1 year ago)
- Default Branch: main
- Last Pushed: 2024-10-24T03:52:18.000Z (over 1 year ago)
- Last Synced: 2024-10-24T15:59:37.416Z (over 1 year ago)
- Topics: kernel-density-estimation, multivariate-analysis
- Language: Python
- Homepage: https://pypi.org/project/mvgkde/
- Size: 319 KB
- Stars: 2
- Watchers: 1
- Forks: 0
- Open Issues: 1
-
Metadata Files:
- Readme: README.md
- Contributing: .github/CONTRIBUTING.md
- License: LICENSE
Awesome Lists containing this project
README
mvgkde
MultiVariate Gaussian Kernel Density Estimator in JAX.
This is a micro-package, containing the single class `MultiVarGaussianKDE` (and
helper function `gaussian_kde`) to estimate the probability density function of
a multivariate dataset using a Gaussian kernel. This package modifies the
`jax.scipy.stats.gaussian_kde` class (which is based on the
`scipy.stats.gaussian_kde` class), but allows for full control over the
covariance matrix of the kernel, even per-dimension bandwidths. See the
Documentation below for more information.
## Installation
[![PyPI version][pypi-version]][pypi-link]
[![PyPI platforms][pypi-platforms]][pypi-link]
```bash
pip install mvgkde
```
## Documentation
[![Actions Status][actions-badge]][actions-link]
For these examples we will use the following imports:
```python
import jax.numpy as jnp
import jax.random as jr
import matplotlib.pyplot as plt
import numpy as np
from mvgkde import MultiVariateGaussianKDE, gaussian_kde # This package
```
And we will generate a dataset to work with:
```python
key = jr.key(0)
dataset = jr.normal(key, (2, 1000))
```
Lastly we will define a plotting function:
```python
# Create a grid of points
(xmin, ymin) = dataset.min(axis=1)
(xmax, ymax) = dataset.max(axis=1)
X, Y = np.mgrid[xmin:xmax:100j, ymin:ymax:100j]
positions = np.vstack([X.ravel(), Y.ravel()])
def plot_kde(kde: MultiVariateGaussianKDE) -> plt.Figure:
# Evaluate the KDE on the grid
Z = np.reshape(kde(positions).T, X.shape)
# Plot the results
fig, ax = plt.subplots()
ax.imshow(np.rot90(Z), cmap=plt.cm.gist_earth_r, extent=[xmin, xmax, ymin, ymax])
ax.plot(dataset[0], dataset[1], "k.", markersize=2)
ax.set(
title="2D Kernel Density Estimation using JAX",
xlabel="X-axis",
xlim=[xmin, xmax],
ylabel="Y-axis",
ylim=[ymin, ymax],
)
return fig
```
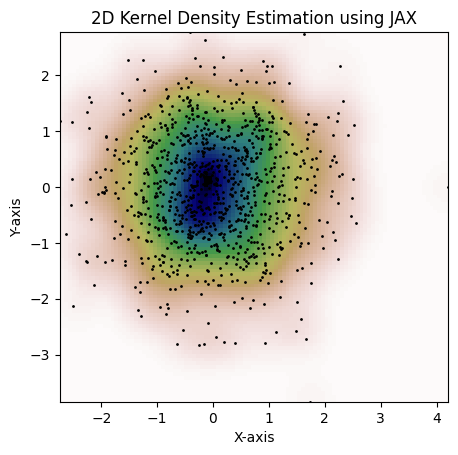
Here's an example that can be done with `jax.scipy.stats.gaussian_kde`:
```python
kde = gaussian_kde(dataset, bw_method="scott")
fig = plot_kde(kde)
plt.show()
```
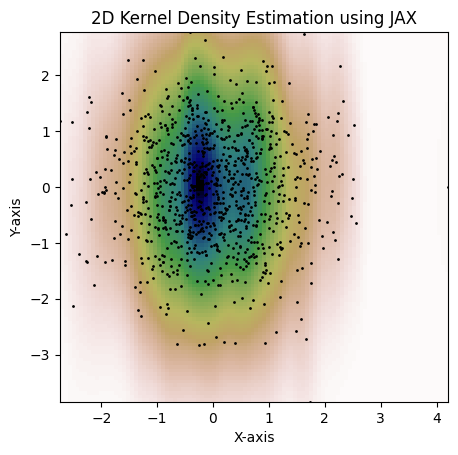
Here's an example with a per-dimension bandwidth. This is not possible with the
`jax.scipy.stats.gaussian_kde`:
```python
kde = gaussian_kde(dataset, bw_method=jnp.array([0.15, 1.3]))
fig = plot_kde(kde)
plt.show()
```
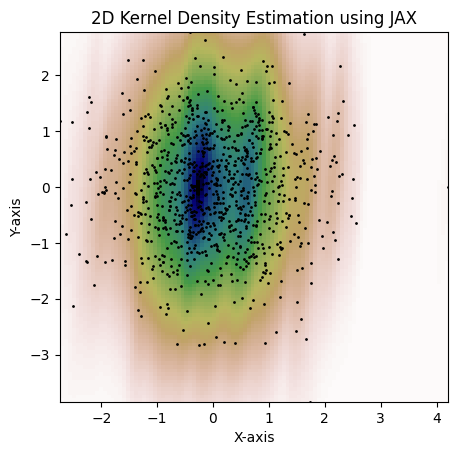
Lastly, here's an example with 2D bandwidth matrix:
```python
bw = jnp.array([[0.15, 3], [3, 1.3]])
kde = gaussian_kde(dataset, bw_method=bw)
fig = plot_kde(kde)
plt.show()
```
The previous examples are using the convenience function `gaussian_kde`. This
actually just calls the constructor method
`MultiVariateGaussianKDE.from_bandwidth`. This function allows for customixing
the bandwidth factor on the data-driven covariance matrix, but does not allow
for specifying the covariance matrix directly. To do that, you can call the
`MultiVariateGaussianKDE` constructor directly, or the `from_covariance`
constructor method. To illustrate the difference between modifying the bandwidth
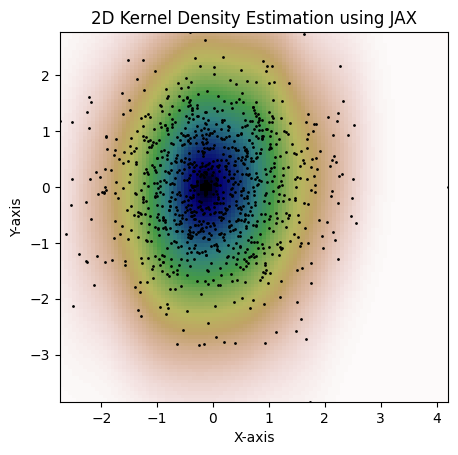
and setting the full covariance matrix, consider the following example:
```python
kde = MultiVariateGaussianKDE.from_covariance(
dataset,
jnp.array([[0.15, 0.1], [0.1, 1.3]]),
)
fig = plot_kde(kde)
plt.show()
```
## Acknowledgments
This package modifies code from [JAX](https://github.com/google/jax), which is
licensed under the Apache License 2.0.
[actions-badge]: https://github.com/nstarman/mvgkde/workflows/CI/badge.svg
[actions-link]: https://github.com/nstarman/mvgkde/actions
[pypi-link]: https://pypi.org/project/mvgkde/
[pypi-platforms]: https://img.shields.io/pypi/pyversions/mvgkde
[pypi-version]: https://img.shields.io/pypi/v/mvgkde