https://github.com/python-graphblas/graphblas-algorithms
Graph algorithms written in GraphBLAS
https://github.com/python-graphblas/graphblas-algorithms
complex-networks graph-algorithms graph-analysis graph-analytics graph-datastructures graph-library graph-theory graph-theory-algorithms graphblas pydata python
Last synced: 2 months ago
JSON representation
Graph algorithms written in GraphBLAS
- Host: GitHub
- URL: https://github.com/python-graphblas/graphblas-algorithms
- Owner: python-graphblas
- License: apache-2.0
- Created: 2022-03-31T04:48:09.000Z (about 4 years ago)
- Default Branch: main
- Last Pushed: 2025-10-06T17:18:07.000Z (7 months ago)
- Last Synced: 2025-10-24T07:44:15.076Z (6 months ago)
- Topics: complex-networks, graph-algorithms, graph-analysis, graph-analytics, graph-datastructures, graph-library, graph-theory, graph-theory-algorithms, graphblas, pydata, python
- Language: Python
- Homepage:
- Size: 391 KB
- Stars: 80
- Watchers: 6
- Forks: 5
- Open Issues: 20
-
Metadata Files:
- Readme: README.md
- License: LICENSE
- Code of conduct: CODE_OF_CONDUCT.md
Awesome Lists containing this project
README

[](https://anaconda.org/conda-forge/graphblas-algorithms)
[](https://pypi.python.org/pypi/graphblas-algorithms/)
[](https://pypi.python.org/pypi/graphblas-algorithms/)
[](https://github.com/python-graphblas/graphblas-algorithms/blob/main/LICENSE)
[](https://github.com/python-graphblas/graphblas-algorithms/actions)
[](https://codecov.io/gh/python-graphblas/graphblas-algorithms)
[](https://doi.org/10.5281/zenodo.7329185)
[](https://discord.com/invite/vur45CbwMz)
`graphblas-algorithms` is a collection of GraphBLAS algorithms written using
[`python-graphblas`](https://python-graphblas.readthedocs.io/en/latest/).
It may be used directly or as an experimental
[backend to NetworkX](https://networkx.org/documentation/stable/reference/classes/index.html#backends).
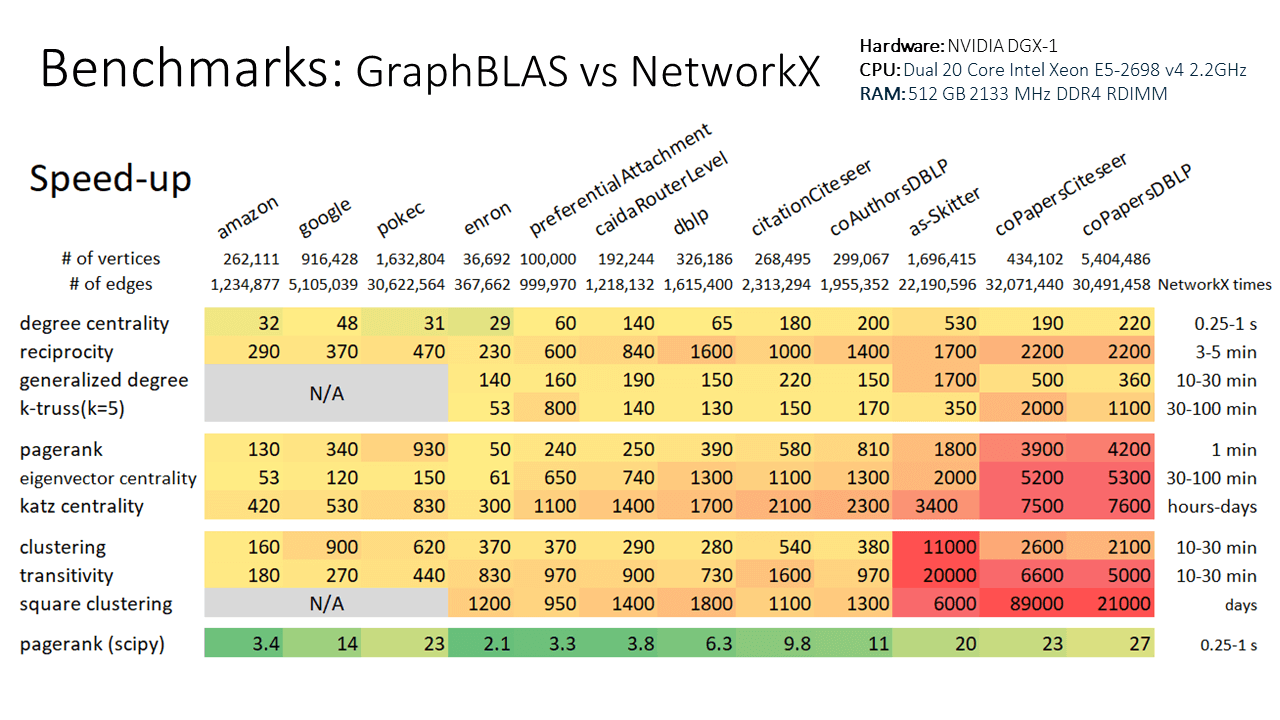
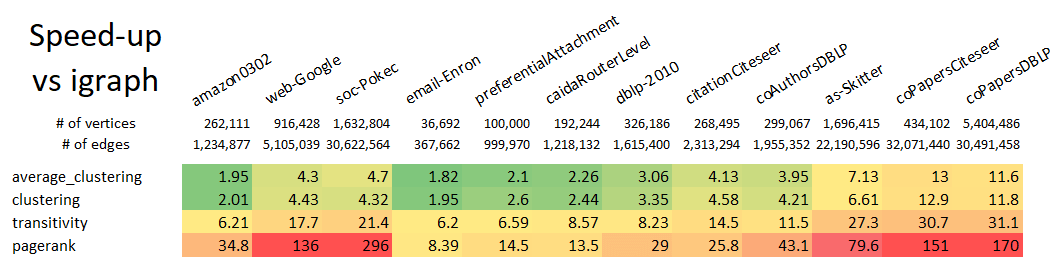
Why use GraphBLAS Algorithms? Because it is *fast*, *flexible*, and *familiar* by using the NetworkX API.
Are we missing any [algorithms](#Plugin-Algorithms) that you want?
[Please let us know!](https://github.com/python-graphblas/graphblas-algorithms/issues)
### Installation
```
conda install -c conda-forge graphblas-algorithms
```
```
pip install graphblas-algorithms
```
## Basic Usage
First, create a GraphBLAS Matrix.
```python
import graphblas as gb
M = gb.Matrix.from_coo(
[0, 0, 1, 2, 2, 3],
[1, 3, 0, 0, 1, 2],
[1., 2., 3., 4., 5., 6.],
nrows=4, ncols=4, dtype='float32'
)
```
Next wrap the Matrix as `ga.Graph`.
```python
import graphblas_algorithms as ga
G = ga.Graph(M)
```
Finally call an algorithm.
```python
hubs, authorities = ga.hits(G)
```
When the result is a value per node, a `gb.Vector` will be returned.
In the case of [HITS](https://en.wikipedia.org/wiki/HITS_algorithm),
two Vectors are returned representing the hubs and authorities values.
Algorithms whose result is a subgraph will return `ga.Graph`.
## Plugin for NetworkX
Dispatching to plugins is a new feature in Networkx 3.0.
When both `networkx` and `graphblas-algorithms` are installed in an
environment, calls to NetworkX algorithms can be dispatched to the
equivalent version in `graphblas-algorithms`.
### Dispatch Example
```python
import networkx as nx
import graphblas_algorithms as ga
# Generate a random graph (5000 nodes, 1_000_000 edges)
G = nx.erdos_renyi_graph(5000, 0.08)
# Explicitly convert to ga.Graph
G2 = ga.Graph.from_networkx(G)
# Pass G2 to NetworkX's k_truss
T5 = nx.k_truss(G2, 5)
```
`G2` is not a `nx.Graph`, but it does have an attribute
`__networkx_plugin__ = "graphblas"`. This tells NetworkX to
dispatch the k_truss call to graphblas-algorithms. This link
connection exists because graphblas-algorithms registers
itself as a "networkx.plugin" entry point.
The result `T5` is a `ga.Graph` representing the 5-truss structure of the
original graph. To convert to a NetworkX Graph, use:
```python
T5.to_networkx()
```
Note that even with the conversions to and from `ga.Graph`, this example still runs 10x
faster than using the native NetworkX k-truss implementation. Speed improvements scale
with graph size, so larger graphs will see an even larger speed-up relative to NetworkX.
### Plugin Algorithms
The following NetworkX algorithms have been implemented
by graphblas-algorithms and can be used following the
dispatch pattern shown above.
[//]: # (Begin auto-generated code)
```
graphblas_algorithms.nxapi
├── boundary
│ ├── edge_boundary
│ └── node_boundary
├── centrality
│ ├── degree_alg
│ │ ├── degree_centrality
│ │ ├── in_degree_centrality
│ │ └── out_degree_centrality
│ ├── eigenvector
│ │ └── eigenvector_centrality
│ └── katz
│ └── katz_centrality
├── cluster
│ ├── average_clustering
│ ├── clustering
│ ├── generalized_degree
│ ├── square_clustering
│ ├── transitivity
│ └── triangles
├── community
│ └── quality
│ ├── inter_community_edges
│ └── intra_community_edges
├── components
│ ├── connected
│ │ ├── is_connected
│ │ └── node_connected_component
│ └── weakly_connected
│ └── is_weakly_connected
├── core
│ └── k_truss
├── cuts
│ ├── boundary_expansion
│ ├── conductance
│ ├── cut_size
│ ├── edge_expansion
│ ├── mixing_expansion
│ ├── node_expansion
│ ├── normalized_cut_size
│ └── volume
├── dag
│ ├── ancestors
│ └── descendants
├── dominating
│ └── is_dominating_set
├── efficiency_measures
│ └── efficiency
├── generators
│ └── ego
│ └── ego_graph
├── isolate
│ ├── is_isolate
│ ├── isolates
│ └── number_of_isolates
├── isomorphism
│ └── isomorph
│ ├── fast_could_be_isomorphic
│ └── faster_could_be_isomorphic
├── linalg
│ ├── bethehessianmatrix
│ │ └── bethe_hessian_matrix
│ ├── graphmatrix
│ │ └── adjacency_matrix
│ ├── laplacianmatrix
│ │ ├── laplacian_matrix
│ │ └── normalized_laplacian_matrix
│ └── modularitymatrix
│ ├── directed_modularity_matrix
│ └── modularity_matrix
├── link_analysis
│ ├── hits_alg
│ │ └── hits
│ └── pagerank_alg
│ ├── google_matrix
│ └── pagerank
├── lowest_common_ancestors
│ └── lowest_common_ancestor
├── operators
│ ├── binary
│ │ ├── compose
│ │ ├── difference
│ │ ├── disjoint_union
│ │ ├── full_join
│ │ ├── intersection
│ │ ├── symmetric_difference
│ │ └── union
│ └── unary
│ ├── complement
│ └── reverse
├── reciprocity
│ ├── overall_reciprocity
│ └── reciprocity
├── regular
│ ├── is_k_regular
│ └── is_regular
├── shortest_paths
│ ├── dense
│ │ ├── floyd_warshall
│ │ ├── floyd_warshall_numpy
│ │ └── floyd_warshall_predecessor_and_distance
│ ├── generic
│ │ └── has_path
│ ├── unweighted
│ │ ├── all_pairs_shortest_path_length
│ │ ├── single_source_shortest_path_length
│ │ └── single_target_shortest_path_length
│ └── weighted
│ ├── all_pairs_bellman_ford_path_length
│ ├── bellman_ford_path
│ ├── bellman_ford_path_length
│ ├── negative_edge_cycle
│ └── single_source_bellman_ford_path_length
├── simple_paths
│ └── is_simple_path
├── smetric
│ └── s_metric
├── structuralholes
│ └── mutual_weight
├── tournament
│ ├── is_tournament
│ ├── score_sequence
│ └── tournament_matrix
├── traversal
│ └── breadth_first_search
│ ├── bfs_layers
│ └── descendants_at_distance
└── triads
└── is_triad
```
[//]: # (End auto-generated code)